Dr. Stedman's Research Lab:
Archaeal Viruses and their Host
This page is being made for a Portland State University Capstone and is a work in progress by Suzie Wallace.
All images have links associated to bring you to the paper they originated as well as the bright orange texts.
All images have links associated to bring you to the paper they originated as well as the bright orange texts.
My Background
I have been working in Dr. Kenneth Stedman's research lab since June of 2019. Before starting here I had taken molecular biology from him where he encouraged everyone to take his Biology C.U.R.E class for Recombinant DNA Laboratory Techniques. In this class I had the opportunity to meet Nichole Procter who was taking this Capstone at the time and was a T.A. for Dr. Stedman's class. She made her own web page explaining C.U.R.E. classes and their purpose. Now, I get to work along side her as I continue to learn how the lab opperates and work on my own projects with her continued support.
My Goals
In the near future I hope to move into graduate school and earn my PhD. My intentions of working in this lab were to understand if research was the path I wanted to take. However, I ended up wanting to go into research by the time the C.U.R.E. class was over. My next goal was to learn how labs regularly operate and get into my own project to keep learning. My main areas of interest are viruses and microbiology. More specifically, I am very interested in Archaea. This lab is the perfect combination of both so my time here has solidified those interests. I have also gained confidence working on an independent project while still having the support and guidance of others.
About the Lab
Dr. Stedman's lab is one of the labs associated with the Center for Life in Extreme Environments at Portland State University. The goal of this lab is to study life where most other organisms can't survive. They are extreme in the sense that they are too hot, too cold, too salty, too acidic, too basic, or under too much pressure. The organisms we work with are single cellular microscopic Archaea and the viruses that infect them. Because they live in extreme circumstances we want to know how they are structured and function to not only live but thrive in these harsh environments.
 Khan Academy, 2019
Khan Academy, 2019
- Outline:
- Virus Background
Sulfolobus Spindle-Shaped Viruses and Sulfolobus - Fun Facts about SSVs and Sulfolobus
Virus Background
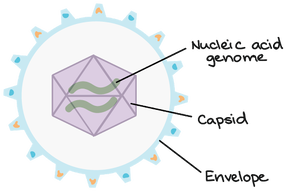
Viruses are nonliving biological entities that do not have the mechanisms necessary to reproduce themselves. They can, however, hijack the mechanisms of living cells to reproduce themselves to go and infect new cells. As seen above, viruses are made of a capsid, nucleic acid, and sometimes an envelope. The nucleic acid can be DNA or RNA and is the genetic material for the virus. The capsid is made of proteins and is used to protect the genetic material as well as interact with the host of the virus. The envelope is a similar structure but is made of fats and proteins that can also protect the genetic material and mediate interactions between the virus and the host. The envelope can even help the virus evade the host's immune response in some cases.
Sulfolobus Spindle Shaped Viruses and Sulfolobus
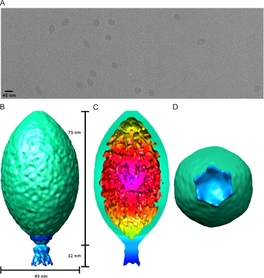 Stedman et. al. Virology, 2015
Stedman et. al. Virology, 2015
Sulfolobus is a genus in the Domain Archaea that includes many species of single celled organisms. These organisms live in hot springs all over the world in temperatures ranging from 75-95°C (167-203 °F) and pH levels ranging from 2-4 (similar to lemon juice!). These cells are hosts for the ever so unique, Sulfolobus Spindle-Shaped Virus. One of the things that makes these viruses so unique is their lemon-shaped capsid. While the virus is not actually these colors, the outside of the virus is split into the green capsid section and the blue tail section on the capsid diagram on the right. The capsid is made of the structural proteins VP1 and VP3 and the tail portion is likely made of the structural protein VP4. On the surface of the host there is a layer of proteins called the S Layer. This is comprised of a single layer of the proteins SlaA and SlaB. The paper I am adapting methods from for my own project is called "Architecture and modular assembly of Sulfolobus S-layers revealed by electron cryo-tomography" . This paper found SlaB to be more of a ruler and a stilt to separate the S Layer from the rest of the cell rather than directly interacting with the virus.
My Favorite Fun Facts about Sulfolobus and SSVs
SSVs systematically and rapidly reproduce in Sulfolobus under UV induction.
 Frols et. al. Virology, 2007
Frols et. al. Virology, 2007
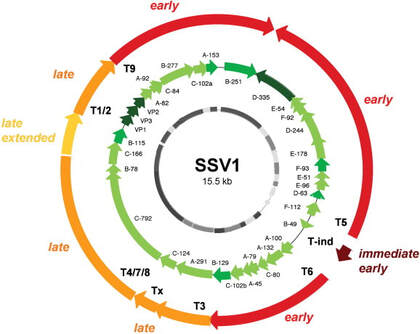
Normally, UV light radiation is damaging to and can kill cells by damaging their genetic material. However, Sulfolobus infected with SSV1 starts to rapidly reproduce the virus after being exposed to UV radiation. The pattern of reproduction is a predictable cycle of up-regulating and down-regulating genes to make proteins at specific time intervals following UV exposure. The image to the left shows this predictable cycle of regulation on the circular genome of the virus. The red to yellow arrows indicate sections of the genes and their timing of regulation where immediate early is right after UV exposure and late is hours after UV exposure. The green arrows are the individual genes that are translated into each protein. When the infected Sulfolobus cells are exposed to UV radiation we can expect to see genes translated into proteins in this predictable order. The exact cause to this pattern is unknown which only makes it more fascinating.
SSVs do not lyse the host cells.
Viruses in general have two primary forms of replication in host cells. The first is the lysogenic cycle where viruses infect the host cell and keep their genes dormant within the hosts cells. That way when the host cell replicates itself and duplicates its' own genetic material, it is also replicating the virus's genetic material and the new cell inherits the infection. The other kind of cycle a virus can have is the lytic cycle. Where the host's molecular replication machinery is actively hijacked to build more viruses. Typically, when the cell has produced enough viruses the cell bursts, or lyses the cell membrane, killing the cell and releasing the newly produced viruses to go on and infect other cells. One of the mysteries of Sulfolobus and some SSVs is that they do not have these relationships. So far, all but two SSVs that infect Sulfolobus have been shown to not lyse the host cell. While viruses are being actively produced the cell does have slowed growth but the host doesn't die. This has raised the question of what, if any, benefit the host cell may be getting from the infection. The virus is clearly getting the benefit of being replicated by the host cell. Maybe the host doesn't benefit from the infection but if it does maybe there is a potentially symbiotic or quasi-symbiotic relationship.
SSVs can withstand a large amount of mutations and still function.
 Goodman & Stedman et al, 2018
Goodman & Stedman et al, 2018
Mutated DNA can mean a variety of things like that the chemical structure of the molecule is damaged in some way or that the original sequence of their genes is altered to some degree. The build up of these mutations is what causes cancer in animals. In this context mutations will refer only to the addition a known large sequence of genes. DNA can be thought of as a track that needs to be shaped a specific way for the machinery to ride down and read to produce RNA and then read again to produce a protein. These insertions into the genetic material disrupt how replication machinery would build the proteins from the genes by changing the message being translated. To better explain how these mutations effect replication the following sentence is how a gene would be read if there was an insertion in the genetic material.
" She went to the zoo and had a fun time lookThere is a parking lot over there.ing at the penguins with her children."
When we read this sentence we can tell that there is a disruption, what both sentences are, and exactly where the insertion is taking place. When a protein is being made from a gene that has been interrupted like this it makes a protein that might not work properly. While we do know what the interrupting sequence is, it takes analysis and preparation to determine where the disrupting gene is in the context of the virus' genome.
The goal of intentionally interrupting their genes is to determine if the proteins made by those genes are essential for the viruses to function. Only a handful of proteins in this genome are defined. There is room to learn about the functionality of any genetic mutation no matter where it is randomly inserted, even if we find the gene is nonessential. We determine the functionality based on the infection and subsequent observation of Sulfolobus growth. Methods used in this process can tell us if these viruses are behaving chemically how we would expect them to, that the mutation is exactly where we determined it to be, and then ultimately if the virus is capable of infecting Sulfolobus with the particular genetic mutation in question.
If a mutant virus is repeatedly observed to not infect the host cells the mutation is considered to be a nonfunctional mutation and we conclude that the protein that was altered by this mutation is essential for virus activity. If the mutation of this gene does not prevent the virus from infecting a cell and replicating itself in the host as usual then we conclude that the protein coded for by this gene is nonessential for virus activity and that the mutation is functional. Interestingly, genes that code for the same protein can function when mutated at one point but can be nonfunctional when mutated at a slightly different point.
If a gene is essential we can then ask, why is this gene essential? Why is this other gene not essential? These questions allow us to learn more about how these mostly undefined viruses operate on a molecular level. SSVs are specifically interesting because they can tolerate almost half of their DNA being mutated and still be infective. With all this room for mutation we can keep making more mutant viruses and continue to learn from them.
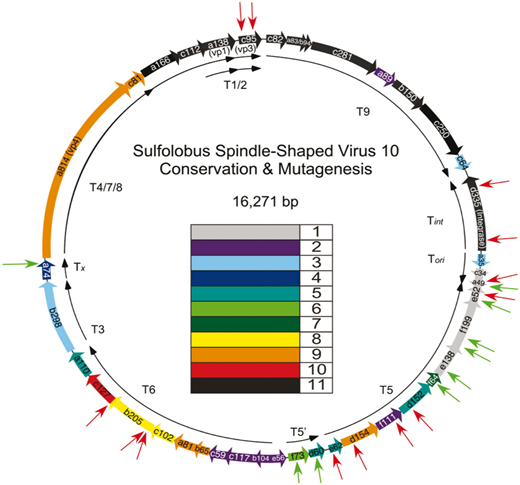
The image above is another circular virus genome, this time of SSV10. The rainbow colored arrows once again represent the individual genes to make each protein. The red arrows pointing at the colored arrows represent previously shown noninfectious mutation insertions and green arrows represent previously shown infectious mutation insertions. The paper this diagram came from is a detailed look at some of these mutations and some of the genes of SSVs. This image is from a paper published by David Goodman and Dr. Stedman. Other papers from this web page were written by past and present members of the lab. SSVs have been mutated in multiple other ways and their applications, analysis of mutants, and process of mutation is simplified for this web page. The papers linked have a much more detailed explanation of all of the topics I have touched on today and further explore the details of Sulfolobus and Sulfolobus Spindle-Shaped Viruses.
Thank you for reading everything and I strongly urge you to continue to read about these viruses and their equally as interesting hosts.